The Three-Dimensional Structure of Nucleic Acids
Nucleic acids, as with any molecule, possess a three-dimensional structure. Nucleic acids can be described into three main levels of structure: the primary, secondary, and tertiary structure. By describing these, we start to unravel why nucleic acids behave the way they do - and why they are capable of being the genetic material found in all forms of life and phages.
The Primary Structure of nucleic acids
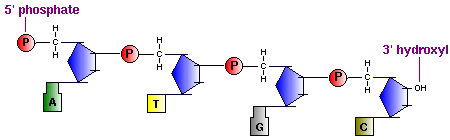
In the diagram above you can see that : the polynucleotide chain has a sense (directionality), where you 'read' it in the direction of the phosphodiester linkage between the 3' carbon of one monomer, to the 5' carbon of the next. The result is the nucleic acid carries an unreacted phosphate (Pi) group on the 5' end, and an unreacted 3' hydroxyl, the 5' to 3' logic is used to 'read' nucleic acids by ribosomes and other proteins involved with nucleic acid synthesis, replication etc. And as is the nature of nucleic acids, they each have a specific nucleotide sequence and for any one nucleic acid, this sequence is its primary structure.
When describing a polynucleotide, it is awkward and quite unnecessary to draw out the entire sequence of nucleotides, the diagram becomes cumbersome. There are several ways of representing the primary structure. For example, if you just want to represent the base sequence, you can do so like this (the first 40 nucleobases of human insulin, on chromosome 11) :
5' ~[... AGCCCTCCAG GACAGGCTGC ATCAGAAGAG GCCATCAAGC ...]~ 3'
In between each letter is a phosphodiester bond, all of these can be assumed to bond to be from a 3' hydroxyl to a 5' phosphate on the next nucleotide. Where the primary sequence ends with a 3', is where the unreacted hydroxyl group is located; conversely, the 5' end is where the unreacted phosphate group occurs.
Hydrogen bonds and helices - the secondary & tertiary structure
The amazing helical shape that nucleic acids can form is familiar with most, it and similar structures occur due to the nucleobases interacting in respect to one another - this interaction results in the secondary structure. Some examples include the tRNA molecule and the double helix found in DNA. The secondary structure of DNA can vary in many forms (conformations) notably the A,B and Z forms. The majority of DNA adopts the B-DNA form, on the other hand, double-stranded RNA and DNA-RNA hybrids usually adopt the A conformation. These forms can occur due to changes in the chemical environment of the cell (including hydration level). You can see them below:
The tertiary structure on the other hand occurs only because of 'longer-range' interactions in the secondary structure. The best example of this occurs when DNA supercoils, compacting it and allowing the large molecule to fit inside cells and phage particles. Supercoiling is present in circular DNA and linear DNA. To understand how supercoiling works, first consider a B-DNA molecule that is a base pairs in length; B-DNA typically has 10.5 base pairs per helical turns. Now that we know this we can consider this B-DNA molecule completes y amount of turns, this number is what we call the Twist (T); if the circular DNA covalently joins at the last turn we say the molecule has a Linking number (L) of y. If this molecule.
If we rotate this molecule counter-clockwise by one turn (360o), it becomes strained (as this reduces the Twist of the helix making it less stable), causing the B-DNA molecule to have 11.67 base pairs per turn (bp/turn). If the strained B-DNA molecule is allowed to return to its more stable conformation, it writhes (W) into a helix with 10.5 bp/turn once again. We can define the writhe as being negatively supercoiled, as W will have a value of -1. If the B-DNA molecule is rotated by two turns, W will have a value of -2, and so on. Overwound DNA is the opposite of this, by rotating it clockwise one turn W will be +1, and so on.

The degree of coiling in supercoiled DNA can be defined by the superhelix density (𝛔), which equals the change in linking number (𝚫L) over the linking number of the relaxed structure (L0), this relationship can be written as:
𝛔 = 𝚫L/L0
Coiling also occurs in single-stranded nucleic acids whether they are RNA or DNA. For example:
- In denatured single strands there is considerable flexibility of nucleotides (residues), resulting in coils and no specific structure.
- Non-self-complementary single strands, such as mRNA create a "stacked-base" structure, where the nucleobases stack, pulling the polynucleotide chain into a non-hydrogen bonding helix. The stacked-base structure is the normal shape assumed by these types of nucleic acid structure under physiological conditions.
- Finally, the "hairpin" structure forms when single stranded nucleic acids are self-complementary. The strand folds back on itself, making a stem-loop structure, much like a hairpin. This can be seen in tRNA, an important molecule in gene expression. See below:

If you'be been reading my previous blogs, you would've noticed that the primary, secondary and tertiary structure of nucleic acids is somewhat analogous in definition to that of proteins. This is because both class of molecule share the same chemical principles when it comes to observing the structure of three-dimensional biological macromolecules.
Further reading:
As always, thanks for reading!
Links provided bring you to some of the info I used, the first year university textbook, "Biochemistry: Concepts and Connections," 1ED, by D.R. Appling, Pearson Ed. LTD, was used as a guide to write this post. You can buy it here.


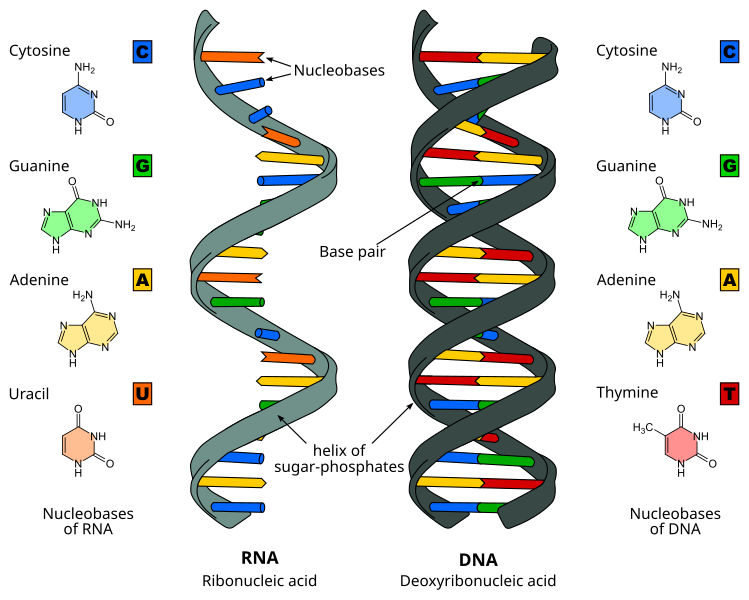
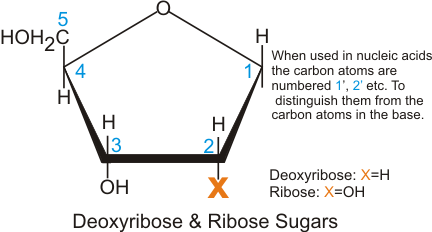


 A consequence of highly conjugated chemical structures is how much the molecules absorb light. Purines, pyrimidines and their derivatives (nucleic acids, nucleotides and nucleosides) all absorb light in the ultraviolet region. You can scroll down on the papers
A consequence of highly conjugated chemical structures is how much the molecules absorb light. Purines, pyrimidines and their derivatives (nucleic acids, nucleotides and nucleosides) all absorb light in the ultraviolet region. You can scroll down on the papers 